| Laboratory of Computational Proteomics Research: Modeling and Simulation of biological systems |
 |
|
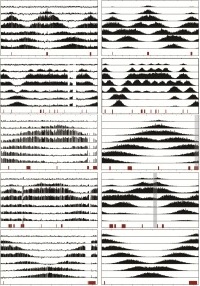
Modeling of replication fork progression throughout the yeast genome
We assayed replication dynamics using whole genome time-resolved chromatin immunoprecipitation combined with microarray analysis of the GINS complex, an integral member of the replication fork. The data show that the replication fork progresses at highly uniform rates regardless of genomic location, revealing that replication fork dynamics in yeast is simpler and more uniform than previously envisaged. We developed a parsimonious model to accurately simulate fork movement throughout the genome. Our model uses a parsimonious set of assumptions: (1) the firing time of a given origin in a population of cells is normally distributed with a mean firing time specific to the origin; (2) the standard deviation of the firing times is constant for all origins in each simulated region; (3) the velocity of progression is the same for all forks (v=1.6 kb/min); and (4) GINS fall off the chromosomes when adjacent forks collide.
Reference
|
 |